Introduction to protein structure prediction

During the last practical the emphasis was on protein sequence
retrieval and anaysis. We will now slowly turn towards protein
structure and focus on what can be deduced on a protein's structure
based on it's sequence. Specifically, we will predict the structure of
a small protein based on its sequence similarity to another protein,
with known structure.
We are going to predict the structure of the alpha-dendrotoxin from
the green mamba snake. This is the toxin contained in the venom of the
green mamba that endangers the prey after a bite. Click here
for more background information on the green mamba.
First, we will extract the toxin sequence from the SWISS-PROT
database. Search for "alpha-dendrotoxin". Click on the required sequence
(it should be the first one listed: IVBI_DENAN (P00980)), and on the
bottom of the net page, right-click on the link 'View entry in raw
text format (no links)', and save the swiss-prot file to a local file
called "venom.swissprot". Also save the sequence in FASTA format
(bottom right) in a file like venom.fasta or similar, for later use.
Secondary structure prediction of alpha-dendrotoxin
As discussed in the lectures, a protein's sequence (primary structure)
can be used as a basis for a prediction of its secondary
structure. The principle of such methods is based on the fact that
different amino acids and amino acid combinations have different
preferences for different types of secondary structure. Alanine, for
example is often found in alpha helices, whereas prolines are known to
destabilise helices. Automated procedures exist that have optimised
prediction algorithms against a databank of proteins with known
structures. One such prediction program is available as an online server:
JPRED secondary structure prediction server. Submit (copy&paste) the venom
sequence (letter code) in the main window and do not forget to
click the checkbox under 4. to omit the PDB search before
hitting the 'Run' button. The server may take some time to complete,
after which the prediction is presented. View the results in HTML format.
You'll notice that the JPRED server first carried out a multiple
sequence alignment before presenting the secondary structure
prediction.
Question:
Why do you think this is? answer.
The prediction is presented near the bottom of the window, in the line
starting with "jpred". A dash (-) stands for unstructured
(i.e. neither helix nor sheet), E stands for extended, or sheet, and H
stnds for helix. As you can see, the serve predicts the protein to
start from the N-terminus with an unstructured loop, followed by two
beta strands and a short helix.
Tertiary structure prediction
Now that we have the sequence of our protein of interest, we need a
suitable template structure of a homologous protein on the basis of
which we can build a model of the venom structure. For this, we visit
protein we're going to use as a template is the bovine (cow)
pancreatic trypsin inhibitor. In the
search field, search for "trypsin inhibitor bovine". Among the search
results (~ 3rd page), select "4PTI", and from the main 4PTI window, select
"Download/Display File". On the Download File menu,
select the corresponding "PDB" format and no compression
(upper left table entry). You should be prompted for a location where
to download the file "4PTI.pdb".
Have a look at the structure by starting rasmol. In a unix window, on
tha command prompt, type:
rasmol 4PTI.pdb
Please note that the commands in bold
print can be easily transferred to the command prompt with
copy-and-paste (select text by dragging the mouse over it with the left mouse
button pressed, and paste by pressing the middle mouse button).
We now see a so-called wireframe
representation of the protein structure: atoms (with different colors
for the different chemical elements: grey for carbon; red for oxygen
and blue for nitrogen) are not shown directly, but the
bonds between atoms are shown as lines. Under "display", also try
other representations such as "sticks", "spacefill", "ball & stick"
and "cartoons". Note that the structure startes with a long,
unstructured loop, followed by a beta-hairpin (a two-stranded
beta-sheet) and ends with a short alpha-helix. Exit rasmol under "file"
-> "exit".
Homology modeling
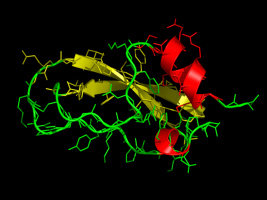
Now we have everything we need to predict a tertiary structure of the
alpha-dendrotoxin from the green mamba snake: its sequence and a
structure of a homologous template. For building the model, we use the
modeling package. To start WHAT IF, type:
/usr/global/whatif/whatif
on the WHAT IF prompt, load the template structure with:
getmol 4PTI.pdb
and press enter if WHAT IF asks for a name.We first need to align the venom sequence that we've retrieved before
with the structure of the bovine trypsin inhibitor structure that
we've just loaded. For this, we first need the protein sequence
corresponding to the protein structure that we have loaded. This can
be done by WHATIF:
%soupir
For residue range, type
all
and as output file name take:
bpti.pir
Now enter the sequence menu:
walign
walseq
First load the sequence that corresponds to the template structure:
getseq bpti.pir
as format, choose
1
Now load the sequence of the green mamba venom:
getseq venom.swissprot
and choose the Swissprot format (3).We now have both sequences loaded and can perform the alignment:
%2align
For the first sequence, choose1
and for the second
2
and choose default values for the gap-open penalty and the
gap-elongation penalty.
We see that the percentage of sequence identity is only 37%. So our
task is now to predict a protein structure based on a structure of
which almost two-thirds of the sequence is different! First, write out
the aligned sequences for later use:
makseq 1 template.pir 1
makseq 2 model.pir 1
And now it is time to build the actual model:
%getpir template.pir
%getpir model.pir
%bldpir
1
2
all
y
Since we chose to use the "Slow but good" version of the structure
prediction module, WHAT IF will take a moment to complete. As soon as
the WHAT IF prompt returns, write out the model structure with:
%makmol
4PTI.pdb
model.pdb
0
all
0
and exit WHATIF
fullstop
View the structure with
rasmol model.pdb
We will now validate our model structure using a protein structure
validation server. This server compares the structure to a database of
known structures and checks if the geometry (bond lengths and angles),
atom contacts etc. are comparable to other protein structures.
MolProbity server. Visit the main page and start. Upload the model
(model.pdb) using the browse button and enter the main page. After the
calculation is finished, press "Continue".
Now we can analyse the results and view e.g. the main Ramachandran plot (click the
"Analyse geometry without all-atom contacts" and after that "Run programs").
Look at the Ramachandran plot, in either kinemage or PDF format.
The Ramachandran depicts the backbone torsion angles
plus contour lines depicting the most favoured regions (as found
for other proteins). As can be seen, all residues are located in the
favoured regions so there are no outliers to worry about.
Also check some of the other options and look for possible anomalies
in the model structure. Note that such tools can be extremely useful
for identifying possible errors in model structures (or in
experimentally determined structures), but that the real hard test for
our model structure is the comparison to its x-ray
structure. Therefore, we will now download the true structure from the
The entry is called 1DTX.pdb. Retrieve it from the server as we did before and download
it to your local account. View the structure with
rasmol 1DTX.pdb
Question:
How good was the secondary structure prediction?
Comparing the model structure with the x-ray structure is easiest with
the two structures superimposed, such that we can compare atom by atom
where the main differences are located. This can be done with the
program g_confrms. This program needs an additional file, the
generation of which would go beyond the scope of this course, which
can be obtained here . Run g_confrms with the
following options:
g_confrms -f1 1DTX.pdb -n1 index.ndx -f2 model.pdb -o fit_whatif_xray.pdb
(select "4" to select the protein backbone for fitting).
g_confrms prints that the overall deviation between the two structures
(measured over all atoms in the protein backbone, so excluding the
side chain atoms) is about 0.1 nm.
Question:
Does that mean that our model is good or is that really a large
deviation? answer.
g_confrms has written a PDB file with both structures superimposed:
rasmol fit_whatif_xray.pdb
To concentrate only on the protein, and remove the ions from the
rasmol view, type in the rasmol comand line:
restrict protein
andcolor chain
The true structure is colored blue, our modelstructure red. As can be seen, the two structures are rather
similar. Especially the backbone structure is rather well predicted by
the model. Some sidechains, however, show larger deviations.
Now, for comparison, we are going to build a model using an
internet server, the SWISS-MODEL
server. Note that this server requires a working E-mail address.
Put your E-mail address in the specified field, provide a name and title,
and paste the sequence of the snake venom in the sequence window (or
use the SWISS-PROT access code: P00980). Before hitting the "Send
request" button, scroll down to the options, and below "Provide your
own templates", upload the structure of bovine pancreatic trypsin
inhibitor (the file 4PTI.pdb that you've dowloaded before from the
Protein Data Bank) as "template file 1". Further below, at the
"Results options", check "Normal mode". Now, submit the request by
hitting the "Send request" button. Depending on the load of the
server, it may take a couple of minutes for the email to arrive with
the coordinates. Actually, you may receive multiple emails, with a
status of your request. The last E-mail should contain the coordinates
in PDB format as an attachment. If you do not receive this E-mail
within a couple of minutes, you may retrieve the coordinates here.
Assuming your model is called "swissmodel.pdb", superimpose the
coordinates to the correct structure:
g_confrms -f1 1DTX.pdb -n1 index.ndx -f2 swissmodel.pdb -o fit_swiss_xray.pdb
Again select "4" when prompted for a group. View the result with:
rasmol fit_swiss_xray.pdb
Also, compare this model with the model generated by WHAT IF:
g_confrms -f1 model.pdb -f2 swissmodel.pdb -o fit_swiss_whatif.pdb
Select "4" twice when prompted for a group. View the result with:
rasmol fit_swiss_whatif.pdb
Question:
How similar/different are the two models? Which of the two models
would you prefer, and why?