Principal components analysis
Last time, we've simulated the dynamics of a small protein. Although a
potein's function is often intimately linked to its conformational
dynamics, it is not always straightforward to extract the functionally
relavant motions from a simulated trajectory. To illustrate this,
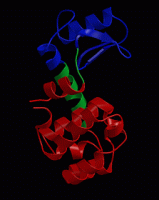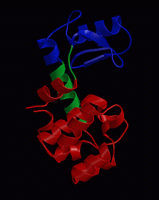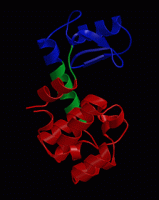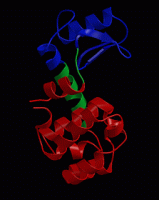
we will focus today on the dynamics of a small, flexible protein:
bacteriophage T4 lysozyme. Many organisms, including humans, have a
form of lysozyme. It degrades polysaccharides (sugar molecules) that
are part of bacterial cell walls and as such it has an antibacterial
function. Click here
or here
for more background information on lysozyme. Download the structure of
T4 lysozyme complexed to a substrat analog (PDB code 148L) from the
Protein Data Bank , and view the
structure in pymol:
/usr/global/whatif/pymol/pymol 148L.pdb
highlight the substrate by typing (on the pymol prompt or in the
grapics window):
select sub,(resn nag,amu)
color red,sub
show sticks,sub
Now select the catalytic residues:
select cat,(resi 11,20,26)
color yellow,cat
show sticks,cat
Now highlight the secondary structure:
dss
show cartoon
As you can see, the substrate is almost enclosed by the protein,
with the active site (catalytic) residues buried in a groove on the
protein surface.
Since it would take to long to calculate a long enough MD simulation
during the course, we have prepared a simulation of the
(substrate-free) enzyme.
Now download this trajectory fragment and view it in pymol:
/usr/global/whatif/pymol/pymol t4l.pdb
Try out different visualisation modes in pymol to view as clearly as
possible the structure and the dynamics of the protein. Use the same
selections as above to highlight the catalytic residues and the
secondary structure and press the "play" button at the bottom right of
the main pymol window to see an animation of the simulation.
Question:
Which motions can you identify? How can you imagine those motions to
be crucial for the protein's function?
Principal components analysis of an MD simulation
Probably, you will appreciate the difficulty of identifying
functionally relevant motions. One of the main reasons for this is
that we see everything moving at the same time. Both local
fluctuations and collective motions occur simultaneously, which makes it
hard to distinguish the two types of motion from each other. A
principal components analysis can help in such cases , as it can
filter global, collective (often slow) motions from local, fast
motions. Download ref.pdb and md1_backbone.xtc. To reduce the size of
the analysis, we will concentrate on the backbone only for the
analysis. First, we have to set up gromacs properly:
source /usr/global/whatif/gromacs/i686-pc-linux-gnu/bin/GMXRC
Now, have a look at the raw (backbone) trajectory:
trjconv -s ref.pdb -f md1_backbone.xtc -o md1.pdb -e 1000
(answer "0" when asked for a group).
/usr/global/whatif/pymol/pymol md1.pdb
and press the "play" button at the bottom right of the main pymol window.
Now start the principal components analysis of this trajectory. This
is done by building a so-called covariance matrix of the atomic
fluctuations. Diagonalisation of this matrix yields a set of
eigenvectors and eigenvalues, that describe collective modes of
fluctuations of the protein. The eigenvectors corresponding to the
largest eigenvalues are called "principal components", as they
represent the largest-amplitude collective motions.
g_covar -s ref.pdb -f md1_backbone.xtc
and answer "0" twice when asked for a group.
If you issue a
ls -lrt
you'll see that g_covar has generated both eigenvalues and
eigenvectors, in files called eigenval.xvg and eigenvec.trr. View the
eigenvalue spectrum with:
xmgrace eigenval.xvg
The eigenvalues are sorted according to their size. As you can see,
there are only a few large eigenvalues, all others are relatively small.
To see what type of motion the indivudual eigenvectors correspond to,
we filter the original trajectory and project out the part along a
selected eigenvector:
g_anaeig -s ref.pdb -f md1_backbone.xtc -filt filter1.pdb -first 1
-last 1 -skip 100
to filter the trajectory along the first eigenvector. View the
animation with:
/usr/global/whatif/pymol/pymol filter1.pdb
Question:
How would you describe this type of motion? How does it compare to the
raw trajectory?
As you can see, the animation is kind of jerky. This is because the
even such large-scale motions do not occur smoothly, but
stochastically. To see a smooth animation of the motion along the
first eigenvector, we can artificially interpolate between the extreme
conformations sampled during the simulaiton along this eigenvector:
g_anaeig -s ref.pdb -f md1_backbone.xtc -extr extreme1.pdb -first 1
-last 1 -nframes 30
View with:
/usr/global/whatif/pymol/pymol extreme1.pdb
Now repeat the last two commads for the second eigenvector.
Question:
How would you describe the motion along the first and second
eigenvector? What happens with the substrate-binding cleft in these
two motions?
To see the contrast with the other eigenvectors, now repeat the above
procedure for eigenvector 20 (or 53, or any other number of your
choice).
Question:
The backbone of T4 lysozyme contains 486 atoms (162 residues and
3 backbone atoms per residue). How many eigenvectors/values would you
expect? How many eigenvalues should be exactly zero, and why? How many
should be approximately zero?
For a more quantitatve analysis, we can project the trajectory onto
individual eigenvectors, and display the projections as a function of
time:
g_anaeig -s ref.pdb -f md1_backbone.xtc -proj
xmgrace -nxy proj.xvg
Question:
Which eigenvector displays the slowest motion? Would you consider the
simulation long enough to have sampled all relevant configurations?
(one criterion for this is that all
sub-conformations should have been visited multiple times).
Another way to visualise the sampled conformations in the subspace
spanned by the eigenvectors is a so-called two-dimensional projection:
g_anaeig -s ref.pdb -f md1_backbone.xtc -2d -first 1 -last 2
xmgrace 2dproj.xvg
In this graph, each point represents a snapshot from the simulation,
and the distribution shows the sampled region along the first two
eigenvectors during the simulation.
Comparison to experimental structural data
We can use a PCA not only to analyse an MD simulation and extract the
main concerted motions from a trajectory, but also for a comparison of
multiple simulations or for the comparison to an ensemble of
experimental structures, in terms of dominant, collective motions.
For T4 lysozyme, a large number of x-ray structures has been determined
(if you search the PDB for "T4 lysozyme", you'll find more than 400
structures). We have collected 38 non-redundant x-ray structures of T4
lysozyme, which are all together collected in allpdb_bb.xtc. Just like on the simulated
trajectory, we can perform a PCA on the collection of experimental
structures:
g_covar -s ref.pdb -f allpdb_bb.xtc
Again, choose "0" twice when asked for a group.
Use the g_anaeig program like above to analyse the results.
Question:
What kind of motion is represented by the first and second
eigenvector? How does this compare to the results from the simulation?
For a quantitative comparison of the simulation and the experimental
strutures, we'll project both the experimental structures and the
simulated structures onto the eigenvectors extracted from the x-ray
structures:
g_anaeig -s ref.pdb -f md1_backbone.xtc -2d md_2d.xvg -first 1 -last 2
g_anaeig -s ref.pdb -f allpdb_bb.xtc -2d xray_2d.xvg -first 1 -last 2
xmgrace md_2d.xvg xray_2d.xvg
You now see the MD simulation in black and the x-ray structures in
red. Since the x-ray structures are independent structures, it is
confusing to have them connected by lines, which suggests a continuous
path between them. In xmgrace, double click on one of the data points
inside the plot, select the second set (G0.S1), under 'line
properties', select 'none', and under 'symbol properties', select
'circle', and press 'Accept'. As you will have seen, eigenvector 1 of
the x-ray ensemble describes an opening motion similar to the
animation on the top-right of this screen, whereas the second
eigenvector describes a twisting motion of both domains with respect
to each other.
Question:
How do the x-ray and MD ensemble compare to each other? Why do you
think the MD simulation does not sample the x-ray configurations on
the extreme left of the plot (the most open x-ray conformations)?
And why do you think the twisting mode is sampled with a larger
amplitude in the simulation than in the ensemble of x-ray structures?
Two additional simulations are available for T4 lysozyme, that started
from different x-ray structures. Download the two backbone
trajectories to your local disk: md2_backbone.xtc and md3_backbone.xtc. Cross project also these
two trajectories onto the x-ray eigenvectors and visualise all three
simulations and the ensemble of x-ray structures projected onto the first
two eigenvectors of the x-ray ensemble.
Question:
Which of the three simulations covers most of the crystallographic
structures? Do you consider the simulations long enough to sample the
most relevant conformations of the protein?
Question:
All three simulations were of the substrate-free protein. Would you
expect a more 'open' or a more 'closed' state as the most probable
configuration in this situation?
Question:
Does the motion along the first eigenvectors appear (quasi)-harmonic?
Which force-constant would you estimate in that case?