Length matters: Hydrophobic mismatch contributes to sorting of SNARE proteins into distinct membrane domains
The view of the lateral organization of lipids and proteins in the plasma membrane has evolved substantially in the last few decades. Nowadays, it is appreciated that most of the plasma membrane proteins and lipids are organized in specific domains. These domains vary widely in size, composition, and stability. They further represent platforms that govern diverse cell functions.

The presynaptic plasma membrane is a well-studied example of a membrane that undergoes rearrangements, especially during exo- and endocytosis. The soluble NSF-attached protein receptor (SNARE) family of proteins has served as an important model for studying the mechanisms that underlie the plasma membrane organization. Initially, cholesterol was shown to be necessary for the functional existence of SNARE domains; however, these domains were different from so-called detergent-resistant membranes (sometimes also referred to as “lipid rafts”)1. Next, the specific protein-protein interactions between cytosolic protein domains were shown to be critical for the segregation of cytoplasmic SNAREs into discreet regions2,3. Interestingly, some of the cytoplasmic SNARE proteins such as syntaxins have a polybasic patch juxtaposed to their transmembrane domain (TMD). Van den Bogaart et al. showed that polyvalent lipids such as phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2) are specifically enriched in certain plasma membrane domains and specifically interact with the polybasic patch of syntaxin 14. Functionally, these syntaxin 1/PI(4,5)P2 domains are shown to act as molecular beacons for vesicle recruitment5.

Figure 3. Syntaxin membrane clustering is induced by a combination of hydrophobic mismatch (increased by cholesterol-induced membrane thickening) and electrostatic interactions with PI(4,5)P2. The membrane clusters are further refined by protein-protein interactions in the aqueous space.
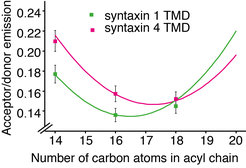
In a collaborative study from our institute, we now show that hydrophobic mismatch suffices to induce clustering of proteins in the plasma membrane6. Hydrophobic mismatch occurs when the TMD of the protein is longer (positive mismatch) or shorter (negative mismatch) than the surrounding lipid environment. The length of the protein TMDs generally increases as we move through the secretory pathway from the endoplasmic reticulum (average length below 20 amino acids) toward the plasma membrane (average length around 27 amino acids)7. However, plasma membrane SNARE syntaxin 1 has a substantially shorter TMD of only 21 amino acids in humans (Fig. 1). Even the crystal structure of the full length SNARE complex reveals that TMDs of syntaxin 1 and synaptobrevin 2 are not long enough to span the plasma membrane, which has an average thickness of around 4 nm7. Indeed, we showed that syntaxin 1 TMD clusters in membranes thinner and thicker than C16:1 phophatidylcholine (PC) (Fig. 2). Molecular dynamics simulations in collaboration with Jelger Risselada and Helmut Grubmüller showed the tendency of syntaxin 1 TMD to oligomerize already at very short time scales (100 µs). The TMD of syntaxin 4 is longer by 1 to 2 amino acids than that of syntaxin 1 and this difference was shown to push the optimal matching (that is, the lowest clustering) toward a thicker bilayer (C18:1 PC). The question arises if these differences in thickness between syntaxin 1 and 4 influence their segregation in the complex lipid environment of the plasma membrane. To this end, we employed two-color superresolution nanoscopy in collaboration with Alf Honigmann, Fabian Göttfert and Stefan W. Hell and observed that, indeed, even this slight difference in thickness (for example 2 amino acids) contributes to segregation of syntaxin isoforms into distinct domains in the plasma membrane. Additionally, since incorporation of cholesterol affects the thickness of the membrane, this study also provides an alternative to the existing view on the cholesterol effect on membrane domain formation. It is important to note that the plasma membrane does not have a uniform thickness, but it rather encompasses a range of thicknesses (between 3.5 and 4.5 nm) depending on the lipids and proteins at the particular domain. Our model proposes that hydrophobic mismatch between lipids and the TMDs of proteins can affect the lateral segregation of particular proteins in the plasma membrane.
All of the mentioned mechanisms (i) protein-protein interactions, (ii) ionic protein-lipid interactions, (iii) cholesterol-induced clustering, and recently described (iv) hydrophobic mismatch-induced clustering act synergistically in establishing the SNARE clusters in the plasma membrane (Figure 3). Furthermore, there is a dynamic exchange of proteins and lipid between these domains. It is tempting to speculate that exactly the formation of the functionally relevant fusion complexes occurs at the interface of these domains.
References
1. Lang T, Bruns D, Wenzel D, Riedel D, Holroyd P, Thiele C, Jahn R: SNAREs are concentrated in cholesterol-dependent clusters that define docking and fusion sites for exocytosis. EMBO J 20, 2202–2213 (2001).
2. Sieber JJ, Willig K I, Heintzmann R, Hell SW, Lang T: The SNARE motif is essential for the formation of syntaxin clusters in the plasma membrane. Biophys J 90, 2843–2851 (2006).
3. Sieber JJ, Willig K I, Kutzner C, Gerding-Reimers C, Harke B, Donnert G, Rammner B, Eggeling C, Hell SW, Grubmüller H, Lang T: Anatomy and dynamics of a supramolecular membrane protein cluster. Science 317, 1072–1076 (2007).
4. van den Bogaart G, Meyenberg K, Risselada HJ, Amin H, Willig KI, Hubrich BE, Dier M, Hell SW, Grubmüller H, Diederichsen U, Jahn R: Membrane protein sequestering by ionic protein-lipid interactions. Nature 479, 552–555 (2011).
5. Honigmann A, van den Bogaart G, Irageta E, Risselada HJ, Milovanovic D, Mueller V, Müllar S, Diederichsen U, Fasshauer D, Grubmüller H, Hell SW, Eggeling C, Kühnel K, Jahn R: Phosphatidylinositol 4,5-bisphosphate clusters act as molecular beacons for vesicle recruitment. Nat Struct Mol Biol 20, 679–686 (2013).
6. Milovanovic D, Honigmann A, Koike S, Göttgert F, Pähler G, Junius M, Müllar S, Diederichsen U, Janshof A, Grubmüller H, Risselada HJ, Eggeling C, Hell SW, van den Bogaart G, Jahn R: Hydrophobic mismatch sorts SNARE proteins into distinct membrane domains. Nat Commun 6, 5984 (2015).
7. Sharpe HJ, Stevens TJ, Munro S: A comprehensive comparison of transmembrane domains reveals organelle-specific properties. Cell 142, 158–169 (2010).
8. Stein A, Weber G, Wahl MC, Jahn R: Helical extension of the neuronal SNARE complex into the membrane. Nature 460, 525–528 (2009).
9. Mitra K, Ubarretxena-Belandia I, Taguchi T, Warren G, Engelman DM: Modulation of the bilayer thickness of exocytic pathway membranes by membrane proteins rather than cholesterol. Proc Natl Acad Sci USA 101, 4083–4088 (2004).





