RIBOFOLD
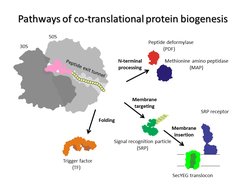
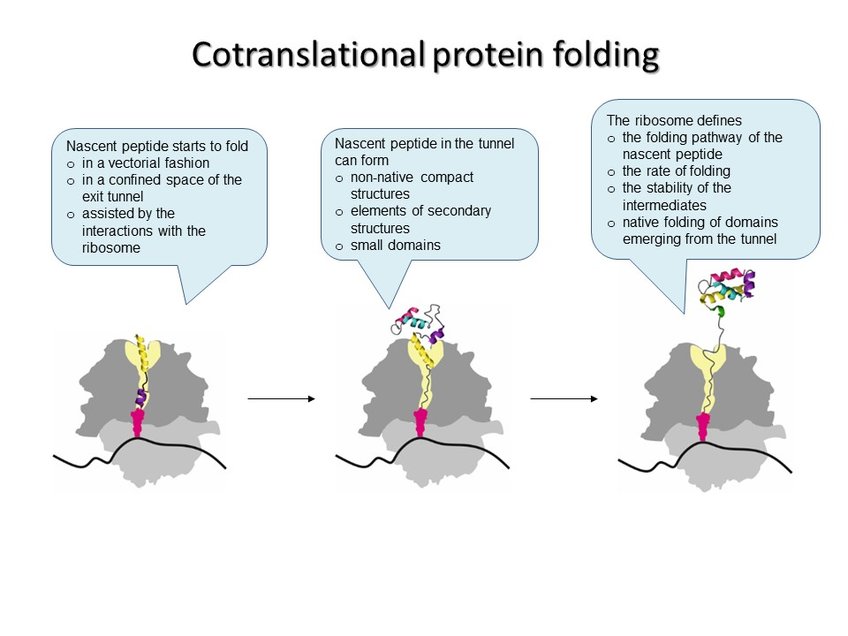
In every living cell, proteins must adopt specific three-dimensional structures to fulfill their diverse functions. Proteins start to fold while they are being assembled by the ribosome during translation. A newly synthesized protein nascent chain enters the exit tunnel of the ribosome, which provides a narrow space that confines protein folding. Each time the ribosome incorporates an amino acid, the growing peptide moves down the exit tunnel until it emerges from the ribosome. When the synthesis is completed, ready proteins are released from the ribosome and can start to execute their cellular functions. Those proteins that happen to be misfolded are re-folded with the help of the chaperones or degraded by the cellular quality control machinery. During its lifetime in the cell a protein can undergo structural rearrangements due to binding to its ligand and partner proteins, but its initial fate depends on the correct structure attained during the pioneering folding event on the ribosome. The vectorial nature of translation, the spatial constraints of the exit tunnel, and the electrostatic properties of the ribosome-nascent peptide complex define the onset of early folding events. The ribosome can facilitate protein compaction, induce the formation of intermediates that are not observed in solution, or delay the onset of folding. Examples of single-domain proteins suggest that early compaction events can define the folding pathway for some types of domain structures. Folding of multi-domain proteins proceeds in a domain-wise fashion, with each domain having its role in stabilizing or destabilizing neighboring domains. The assembly of protein complexes can also begin cotranslationally. In all these cases, the ribosome helps the nascent protein to attain a native fold and avoid the kinetic traps of misfolding. Finally, the pace of translation can change protein folding to produce more or less of the functional peptide form.
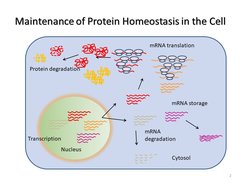
Defects in protein folding disturbs the cellular proteostasis, which can result in debilitating diseases. Single amino-acid substitutions can disrupt a protein’s structure in the cell to cause, for instance, cystic fibrosis, sickle cell anemia, cataract, Huntington’s disease, or retinitis pigmentosa. The molecular pathology of these diseases is a perturbation of the native three-dimensional structure leading to a misfolded protein that can no longer execute its function and is prone to aggregation and rapid degradation. Furthermore, mutations in natively disordered proteins, such as α-synuclein, tau protein or amyloid β-peptide, can cause aggregopathies, such as Parkinson’s and Alzheimer’s.
Despite its importance for understanding human diseases, the mechanisms of co-translational folding and the link between the speed of translation and the quality of protein folding are poorly understood. The aim of the RIBOFOLD project is to understand when, where and how proteins emerging from the ribosome start to fold, how the ribosome and auxiliary proteins bound at the polypeptide exit affect nascent peptide folding, what causes ribosome pausing during translation, and how pausing affects nascent peptide folding. To address these questions, we utilize a toolbox of ensemble and single molecule biophysical techniques to monitor translation and protein folding simultaneously at high temporal resolution. We expect that these results will open new horizons in understanding co-translational folding and help understand the molecular basis of many diseases.